Research Aim
Humans and other animals can retain knowledge gained from daily experience and learning throughout their lifetime. The question of where and how such memories are stored in the body has fascinated people since ancient times. Today it is widely accepted that memory is stored in the brain — yet the details of which regions store memories and by what mechanisms remain largely unknown.
The goal of our laboratory is to elucidate, at the cellular and molecular level, where, when, and how memories are stored in the brain, and how they are recalled. Using laboratory animals, we combine state-of-the-art techniques from molecular biology, biochemistry, physiology, optics, and behavioral science to reveal how the properties of cortical neurons and neural circuits change during the formation and maintenance of memory.
Current Research Topics
1. Functional analysis of the activity-dependent gene Arc [Okuno]
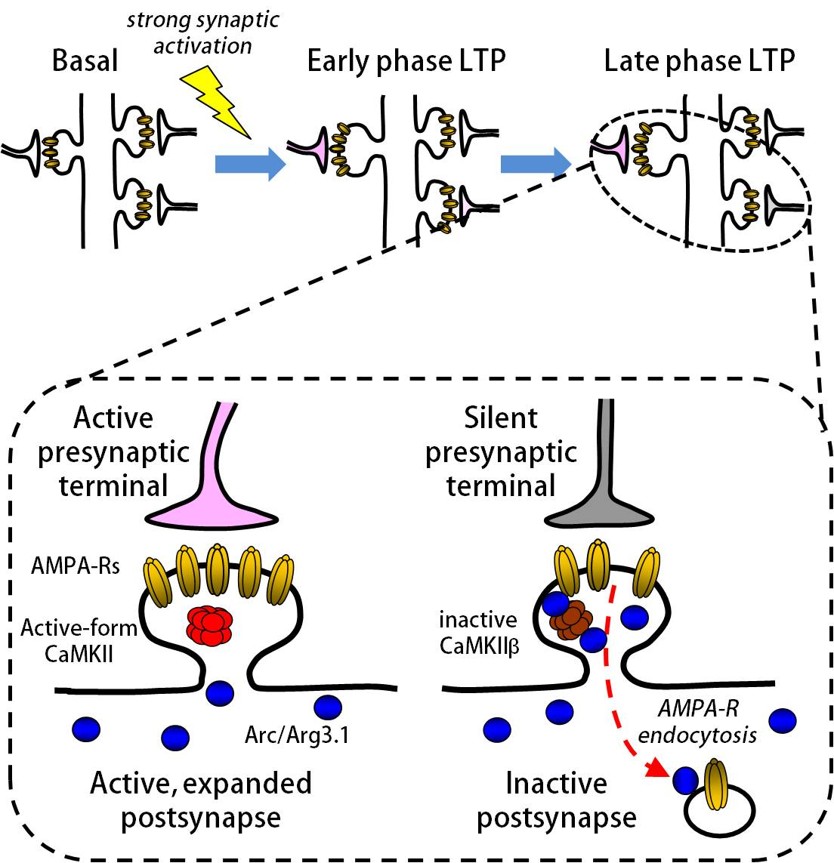
The activity-dependent gene Arc (Activity-regulated cytoskeletal-associated protein, also known as Arg3.1) is known to regulate synaptic strength. We have previously shown that Arc protein is recruited to postsynaptic sites through binding to the β subunit of CaMKII, and that this regulates the synaptic surface expression of glutamate receptors (Chowdhury et al., Neuron, 2006; Okuno et al., Cell, 2012). Genetic studies in mice have shown that the Arc gene plays a critical role in long-term memory, but the detailed mechanism remains unclear. We are currently investigating where and when Arc expression is required for memory formation and maintenance.
2. Visualization and identification of memory-related cortical circuits [Okuno, Kiyama]
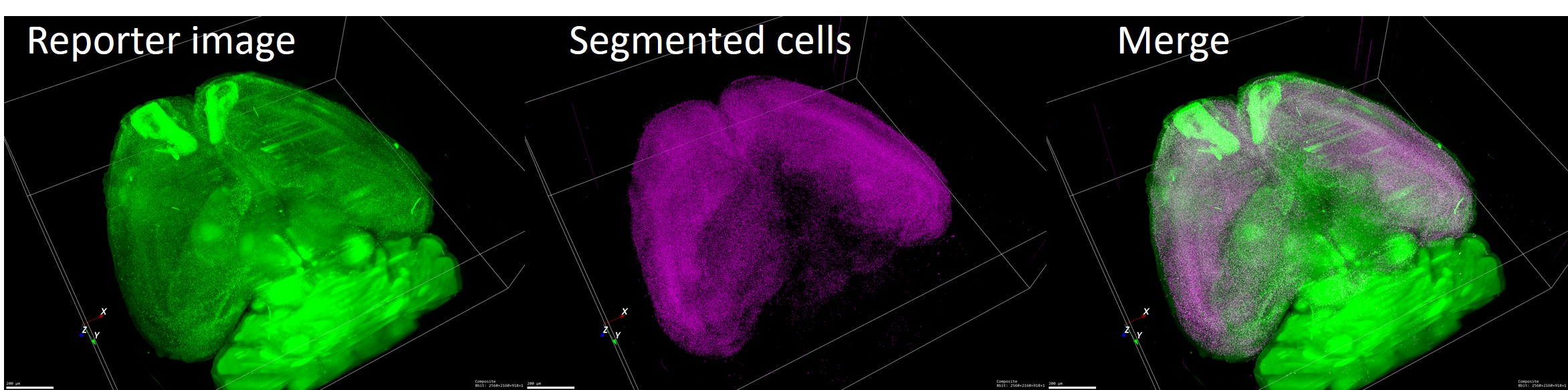
Memory formation does not engage every neuron in the cortex; rather, particular neuronal ensembles are recruited as a memory circuit for a given memory. By visualizing these neuronal ensembles, we aim to determine where in the brain memories are formed and maintained. We have developed several activity-dependent fluorescent reporter systems based on activity-induced promoters (Kawashima et al., PNAS, 2009; Kawashima et al., Nat Methods, 2013; Mikuni et al., Neuron, 2013; Vousden et al., Brain Struct. Funct., 2014). We are now generating activity-reporter mice based on these technologies and combining them with tissue-clearing methods to identify memory-related neural circuits comprehensively at the whole-brain scale.
3. Identification of novel synaptic-function regulating genes [Okuno, Haraguchi]
In addition to Arc, many genes — including c-fos, fosB, egr-1 (zif268 / NGFI-A), and Nr4a1 (NGFI-B / Nur77) — are transiently induced by neuronal activity. However, with a few exceptions, the analysis of these genes is far from complete and the functions of many remain unknown. By investigating activity-dependent genes that have not yet been studied in detail, we aim to discover new regulators of synaptic function.
4. Development of new cognitive tasks [Kiyama, Suzuki]
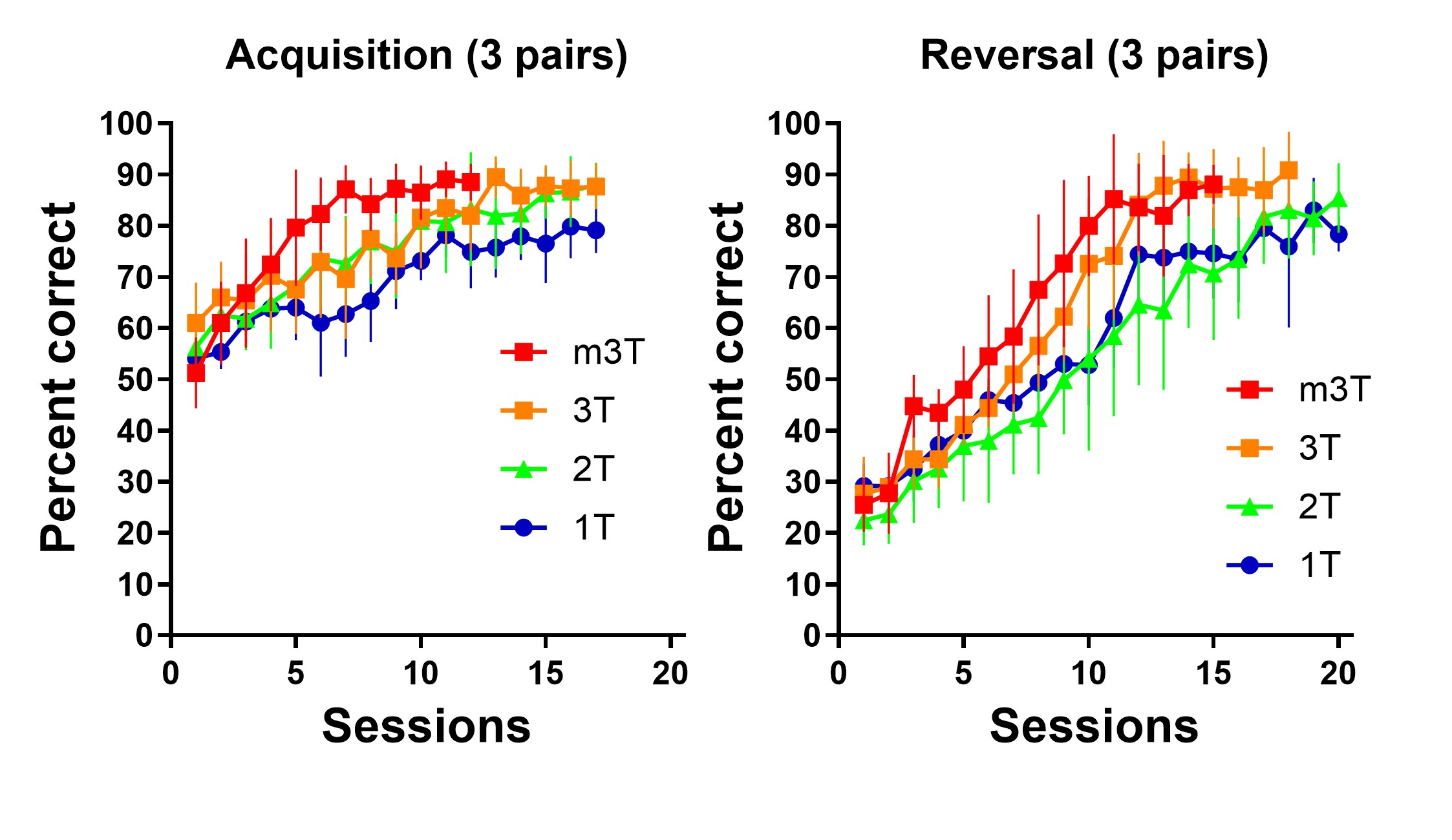
To investigate cognitive functions including learning and memory in laboratory animals such as mice, behavioral tasks suited to each purpose are essential. In addition to classical tasks for testing memory and emotion, we are using touchscreen-based operant systems to develop novel behavioral tasks and learning protocols.
5. Functional analysis of the cortico-thalamic network [Kuramoto]
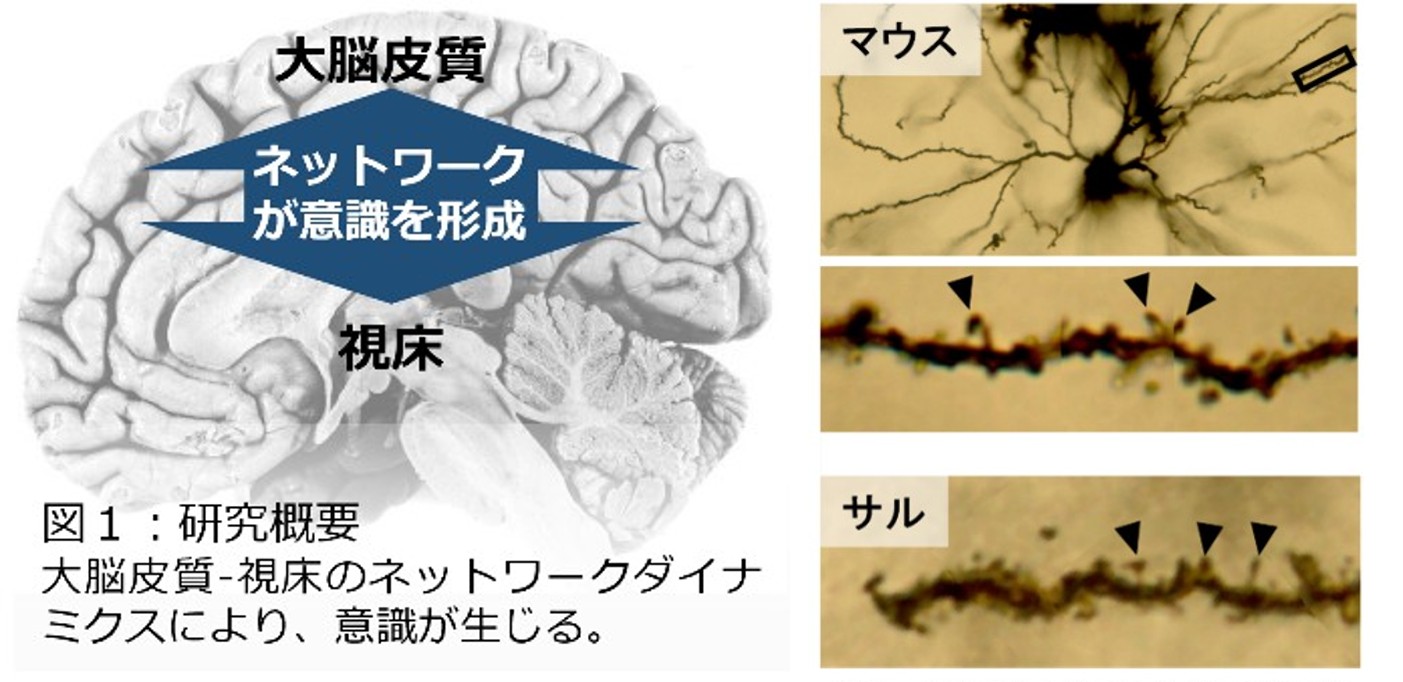
The thalamus has long been known to relay information from the external world to the cerebral cortex. However, large-scale lesions of the thalamus cause loss of consciousness even when the cortex itself is intact. This suggests that consciousness arises not from the cortex alone but from the cortico-thalamic network as a whole, a hypothesis we aim to test. Recently we have found that thalamic neurons also possess dendritic spines, and we are now characterizing their function.
Selected References
- Chowdhury S, Shepherd JD, Okuno H, et al. Arc interacts with the endocytic machinery to regulate AMPA receptor trafficking. Neuron, 42: 445–459 (2006).
- Kawashima T, Okuno H, et al. A synaptic activity-responsive element in the Arc/Arg3.1 promoter essential for synapse-to-nucleus signaling in activated neurons. Proc. Natl. Acad. Sci. USA, 106: 316–321 (2009).
- Okuno H. Regulation and function of immediate-early genes in the brain: beyond neuronal activity markers. Neuroscience Res. 69: 175–186 (2011).
- Okuno H, Akashi K, Ishii Y, et al. Inverse synaptic tagging of inactive synapses via dynamic interaction of Arc/Arg3.1 with CaMKIIβ. Cell, 149: 886–898 (2012).
- Kawashima T, Kitamura K, Suzuki K, et al. Functional labeling of neurons and their projections using the synthetic activity-dependent promoter E-SARE. Nat Methods, 10: 889–895 (2013).
- Kiyama Y, Suzuki Y, Haraguchi M, et al. A simple method that doubles learning speed for mice in touchscreen-based visual discrimination. Sci Rep, 15: 43133 (2025).